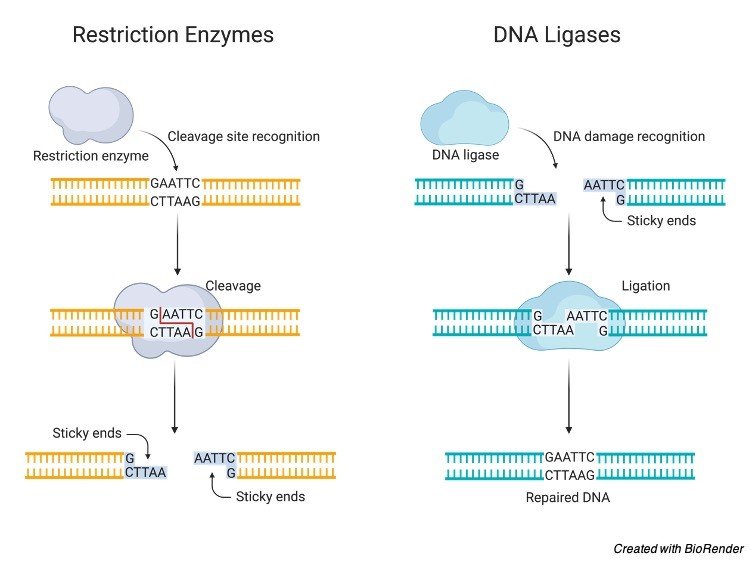
Type IIS restriction enzymes cleave outside of their recognition sequence. For a comprehensive review of restriction endonucleases, see Fuchs and Blakesley. Golden Gate assembly utilizes a Type IIS restriction enzyme to generate DNA fragments with compatible overhang sequences, and a DNA ligase to join the fragments together. The cleavage sites fall into three different categories, either flush (or blunt) in which the recognition site is cut in the middle, or either with 5' or 3' overhangs, in which case unpaired bases will be produced on both ends of the fragment. Each enzyme cuts the palindrome at a particular site, and two different enzymes may have the same recognition sequence, but cleave the DNA at different points within that sequence. The human genome has many palindromic sequences distributed throughout. Palindromic sequences are found in abundance in the genome of most organisms. The recognition site for most of the commonly used enzymes is a short palindromic sequence, usually of either 4, 5, or 6 bp in length, such as AGCT (for Alu I), GAATTC (for Eco RI), and so on. Most restriction enzyme recognition sites are palindromic and include only specified base pairs (i.e., EcoRI recognizes GAATTC). The recognition sequence of few restriction enzymes is given below: DNA Replication, Gene Regulation and Expression Palindromic sequences have great significance. This has to be determined for each individual enzyme. The name of the enzyme (such as Bam HI, Eco RI, and so forth) tells us about the origin of the enzyme, but does not give us any information about the specificity of cleavage. The nomenclature of enzymes is based on a simple system, proposed by Smith and Nathans. These enzymes can be purchased from the many manufacturers of biotechnology products. Cleavage is at the recognition site (but may occasionally be just. Restriction endonucleases are bacterial enzymes that cleave duplex DNA at specific target sequences with the production of defined fragments. Site recognition is via short palindromic base sequences that are 4-6 base pairs long. The ability to cleave DNA at specific sites is one of the cornerstones of today's methods of DNA manipulation.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
